Leonid A. Mirny
Associate Member, Broad Institute at Harvard and MIT
Research Interests
The challenge of understanding biological systems from first physical principles is what motivates our research. We believe that simple physical models are valuable for describing these systems.
- Physics of Chromosomes – exceedingly long polymers that carry genetic and epigenetic information. We aim to understand the role of dynamics and 3D organization of chromosomes in regulation of gene expression, execution of genetic programs, DNA repair and compaction.
- Polymer Physics
- Active Polymer systems
- Epigenetic (neuronless) memory
Broadly, our goal is to understand how physical mechanisms underlie reading, writing and transmission of genetic and epigenetic information. A key feature of our approach are the direct collaborations we have with other scientists in the area. A number of graduate students in the lab are co-advised by an experimentalist. Nearly every student in the lab works directly with our experimental collaborators, contributing both to experimental design, data analysis, and modeling.
Courtesy of Serious Science | YouTube
Biographical Sketch
Leonid Mirny combines biophysical modeling with analysis of large genomics data to address fundamental problems in biology. Mirny aims to understand how long DNA molecules of chromosomes are folded in 3D, and how this 3D organization is used by living cells. The Mirny group has proposed that DNA is folded by molecular motors that perform “loop extrusion”. The loop extrusion hypothesis has been confirmed experimentally and has revolutionized our understanding of chromosomes across all organisms. Now Mirny aims to understand how cells use this and other physical mechanisms to regulate its genes, repair DNA, and maintain epigenetic memory.
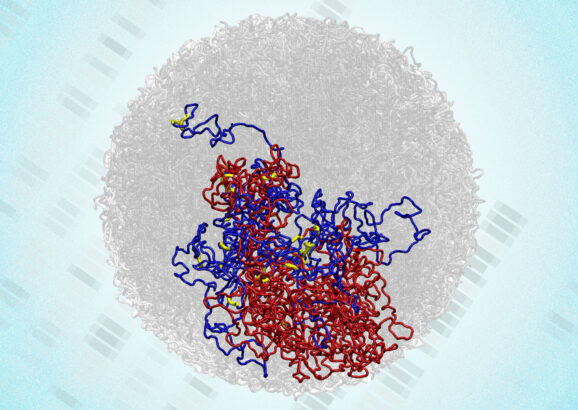
In a surprising discovery, scientists find tiny loops in the genomes of dividing cells
Enabled by a new high-resolution mapping technique, the findings overturn a long-held belief that the genome loses its 3D structure when cells divide.
Awards & Honors
- 2024 // Tel Aviv International Prize in Biophysics
- 2023 // Simons Investigator
- 2019 // Blaise Pascal International Chair of Excellence
- 2014 // American Physical Society Fellow "for elucidating principles of protein-DNA search, and for applying concepts and methods of polymer physics to characterize the three dimensional organization of genome within a cell."
- 2003 // Alfred Sloan Research Fellow
- 2003 // NEC Fund Award
- 2001 // John F. & Virginia B. Taplin Award
- 1999 // William F. Milton Award
